S. marcescens nuclease
S. marcescens endonuclease, also known under commercial name Benzonase®, is extracellularly secreted enzyme of Serratia marcescens, which is supposed to be one of the virulence factors contributing to pathogenicity of this bacterium. However, this enzyme has been widely employed within a number of techniques in biotechnological industry.
Serratia marcescens nuclease is highly effective nonspecific nuclease with very little sequence selectivity digesting both single and double stranded DNA as well as RNA. It has virtually no proteolytic activity making it an ideal candidate for removal of nucleic acids from protein samples, or reducing the viscosity caused by high molecular weight DNA release during cell lysis.
Molecular structure of S. marcescens endonuclease
S. marcescens endonuclease occurs as monomer or dimer both of which are catalytically active, but each subunit cleaves substrate independently. Moreover, these two forms significantly differ in the working mechanism: monomer follows distributive manner, while dimer follows processive manner. However, the dimeric form is more efficient due to higher activity during association with nucleic acids.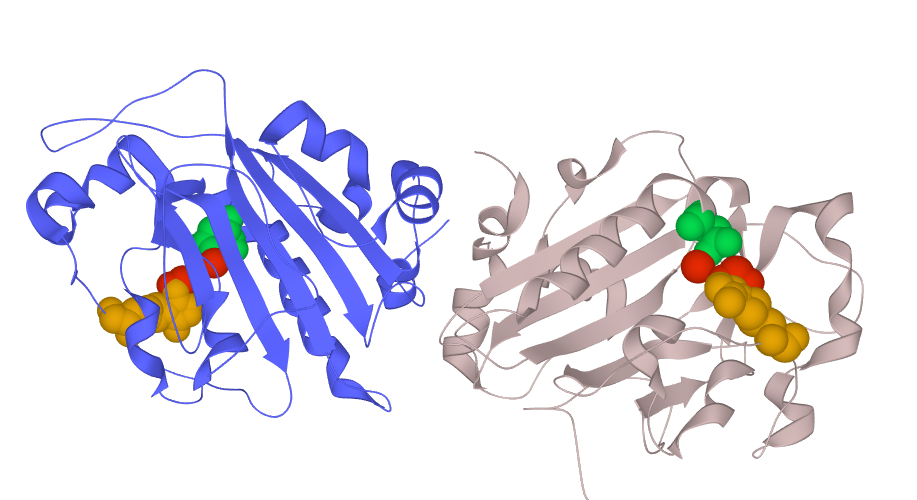
Fig. 3D structure of the S. marcescens nuclease dimer with highlited active sites
Salt requirements of S. marcescens endonuclease
S. marcescens endonuclease shows a strict dependence on divalent cations, especially on Mg2+. Among other ions, which can substitute magnesium, there are Mn2+, Co2+, Ni2+, Zn2+, and Cd2+. Magnesium binding site is located within an active site of each monomer and forms magnesium-water cluster, which structure is conserved within various nucleases homologous to S. marcescens. endonuclease.
Enzymatic mechanism of S. marcescens endonuclease
This enzyme is a nonspecific nuclease, which shows preferences for A-form of double stranded DNA. The active site of this nuclease is characterized by three conserved residues - Arg57, His89 and Asn119. His89 acts as a general base activating the water molecule for a nucleophilic attack on the phosphodiester bond being cleaved. The water molecule forms a hydration sphere of the Mg2+ and protonates the leaving group. Magnesium as a co-factor directly contacts the oxygen of Asn119. Arg57 acts in stabilization of the transition state as well as the leaving group.
Application
• Degradation of DNA after cell lysis
• Viscosity reduction of a sample caused by HMW nucleic acids
• DNA and RNA removal from protein samples
• Replacement of sample sonication or other mechanical shearing of DNA
• Prevention of cell clumping
Activity of Protean’s S. marcescens endonuclease
One unit of this enzyme digests the amount of DNA that changes an A260 of 1.0 in 30 minutes at 37°C under standard assay conditions: 50 mM Tris-HCl (pH 8.0), 1 mM MgCl2, 100 ug/ml BSA, and 1 mg/ml of sonicated salmon sperm DNA.
References:
The Active Site of Serratia Endonuclease Contains a Conserved Magnesium-Water Cluster.
Advantage of Being a Dimer for Serratia marcescens Endonuclease.
A similar active site for non-specific and specific endonucleases.



